M.S. Candidate: Cansu Dinçer
Program: Bioinformatics
Date: 06.09.2019 / 11:00
Place: A-212
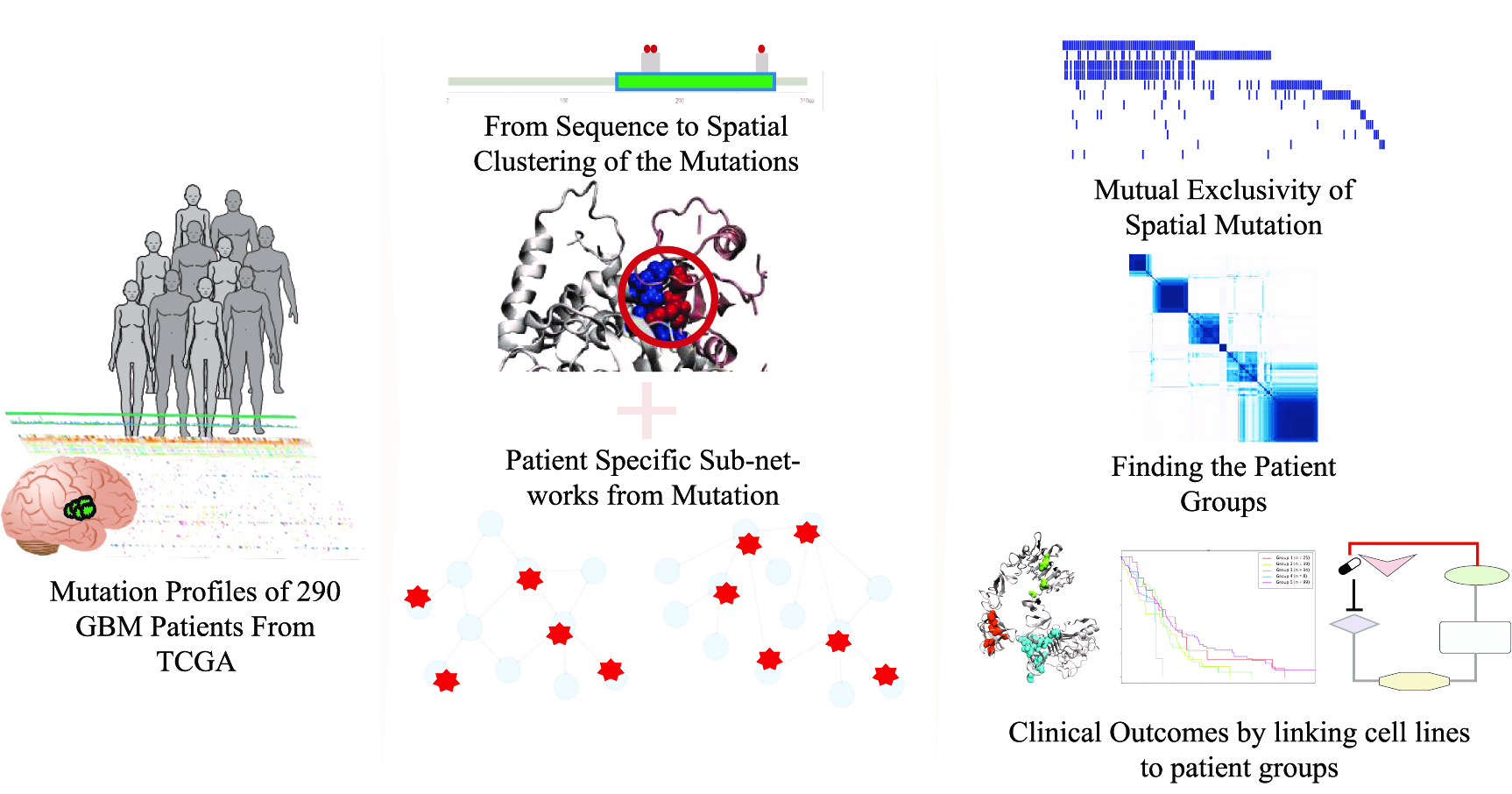
Abstract: Glioblastoma multiforme (GBM) is the most aggressive and heterogeneous type of brain tumor. The heterogeneity of GBM is the main obstacle to develop effective treatment strategies. In this study, we aimed to decrease the heterogeneity among GBM patients from The Cancer Genome Atlas (TCGA), classify the patients and propose therapeutic hypothesis for patient groups by using patient mutation profiles. We therefore implemented a systems level approach to mutations using their biophysical characteristics and organization in patient-specific subnetworks. While 3D mutation patches decrease the heterogeneity among patients, network guided analysis classified patients into five groups. Since each patient group carries a set of signature 3D mutation patches, we collected GBM cell line mutation and drug sensitivity data to link GBM patient group to GBM cell lines and eventually drug responses through 3D patch mutations. Therefore, we can propose drug responses of specific patient group for specific drugs. As an example, by targeting CSF1R, Pazopanib can be effective for Group 3 yet Group 2 can be resistant to inhibition of ATM which is a mediator of PTEN phosphorylation. As a conclusion, from mutations to protein interaction networks and eventually to therapeutic data, this study is a new perspective for precision medicine.